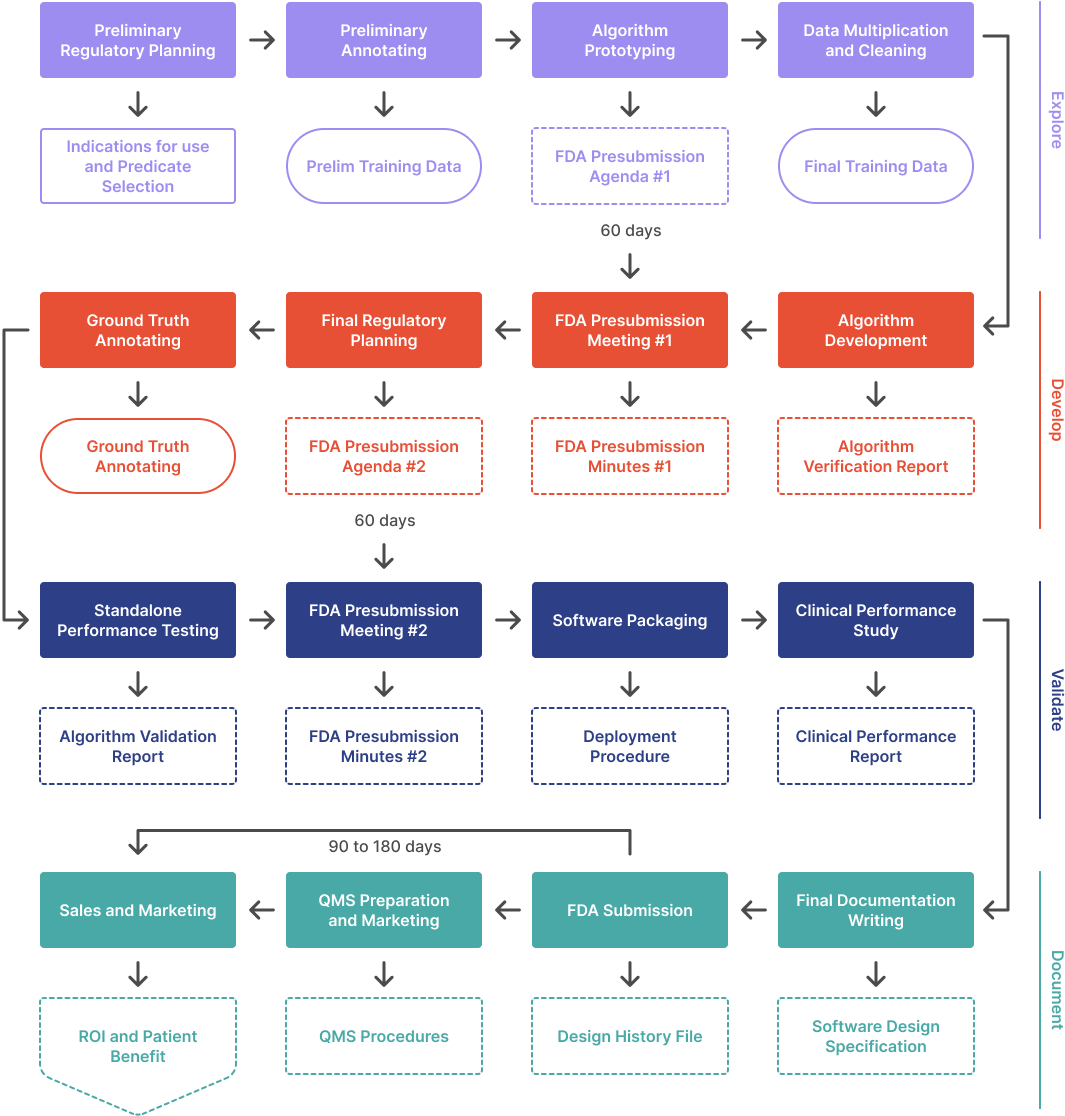
This questionnaire is for you if you already built an AI algorithm built and are looking to get it FDA cleared. It will help set the groundwork for subsequent conversations with FDA, regulatory consultants, or both. Going through this exercise could help you uncover gaps that you may need to address before FDA submission. One of the biggest hurdles to achieving FDA clearance is the validation study. There are two types of validation studies: standalone performance testing and clinical performance testing. In this article, we will review the simpler of the two—standalone performance testing.
For the Device Under Consideration 🔗
- Starting from the input (e.g., a CT scan) and ending at the output (e.g., a report with lung nodules), split the algorithm into smaller software items. Aim for at least 4 but no more than 10.
- Any machine learning model should be its own software item
- If you only have one large machine learning software item, consider breaking it up to add more transparency. FDA does not like big black boxes.
- Traditional image processing steps should be their own module (e.g., morphologic operations, post processing U-net output, etc)
- DICOM loading and report generation should be a software item but should not count towards the limits.
- Have you set aside any labeled data that has not been used by the engineering team to iterate on algorithm development? If so, describe where the data came from.
- Is it possible for you to acquire race, gender, and age? Gender and age are typically found in the DICOM header but race can be harder to come by unless you ask your data broker for this information.
- Describe acceptance criteria for the device as a whole. Even though you will be running the tests yourself, imagine yourself as a skeptical clinician trying to determine if the device works as intended. What acceptance criteria would you propose, such that if they passed, you would be comfortable using it on your patients? For instance, a correlation coefficient of at least 0.99 between the measured tumor volume and the predicted tumor volume would prove the device is fulfilling the clinical use case.
For Each Non-ML Software Item 🔗
- Describe inputs and outputs. Include data shapes, data types, and the transformations undertaken.
- Break the algorithm down into a recipe such that another practitioner could replicate. Think of it as what you would need to document for a patent application. The burden of proof is lower the more you can fully describe how a software item works.
- Devise one test case with synthetic data input and output that is either generated or feasibly double checked manually to be expected. For example, suppose you have a software item named “segmentation cleaner”. The job of this item is to close holes and remove islands smaller than 10 square pixels. You can easily create a synthetic input image with holes and islands and a manually generated output image with the expected result. Usually only one or two such test cases are necessary because these algorithms and their edge cases are well understood.
- If the test cases seem too trivial, consider combining the software item with a proceeding or subsequent step as appropriate.
For Each ML Software Item 🔗
- Describe the input and the output of the machine learning algorithm. I find it helps to include the data type and shape. For example, a cell segmentation algorithm inputs an RGB image of
512x512x3 Float32and outputs a labelled image of512x512x1 uint8where each value is an id for an individual cell. - Describe the data that was used to train the algorithm.
- Where did you source the data from?
- How many images from each site?
- Describe the data annotation protocol.
- Who annotated the data?
- How did you achieve consistency between different annotators?
- Do you have a training manual with examples and how to handle ambiguous situations?
- What qualifications do the annotators hold?
- Describe how you partitioned the training and tuning set. If you are a data scientist, the “tuning” set is the “validation” set. However, the term “validation” set is discouraged because it can be confused with clinical “validation”
- What mechanisms have you used to guard against data leakage between the aforementioned partitions? For example, a common way to do this is to keep data from different institutions in different data partitions. At the very least you should take care the same patient is not in multiple partitions.
- Describe the algorithm architecture (e.g., U-net, transformers, VGG, etc)
- Describe the loss function
- Describe tools you used to detect and prevent overfitting
- Data augmentation
- Cross validation
- Early stopping
- Regularization techniques
- Describe your acceptance criteria for the specific item. Imagine you are outsourcing the algorithm development to an external firm. What acceptance criteria would you add to the statement of work to ensure the quality of the deliverable meets your expectations? For example: Dice score > 0.95
Example 🔗
Here is an example of a hypothetical device that detects malignant lung nodules.
| Software Item Name | Description | Input | Output | Acceptance Criteria | Data Provenance |
|---|---|---|---|---|---|
| DICOM Receiver | A DIMSE compliant C-STORE SCP | DICOM DIMSE Commands | A directory with DICOM files in a patient / study / series directory structure | Passing tests. Manually pushing DICOMs to the | N/A |
| DICOM Loader | Loads 2D DICOM slices into a 3D Nifti volume representation | Previous output | The normalized NIfTI file. Normalized to voxel size of 1mm x 1mm x 1mm in a LAS patient orientation and Hounsfield units. Data type: Float32. Shape: Same as original DICOM | Passing unit tests. | N/A |
| Lung Segmentation | U-net that segments the lung in a CT scan | Previous output | A NIfTI file with the same shape and coordinate system as the input. Value of True in the Lung, False outside lung. |
Dice score of 0.98 | 1000 CT scans. Manually annotated by technicians. Reviewed by board certified thoracic radiologist. |
| Lung Segmentation Cleaner ML | Cleans artifacts of lung segmentation. Removes any islands smaller than 500 cubic millimeters using scipy.label. Fills holes using morphological operations. |
Previous output | A cleaned NIfTI file with | Passing unit tests. Synthetic input and expected output datasets. | N/A |
| Image Filter | Rejects images without the entire lung visible | Previous output |
True if at least 1000 mm3 of lung volume is detected |
Passing unit tests | N/A |
| Lung Nodule Candidate Generator ML | Mask R-CNN algorithm that detects lung nodules | Previous output | Zero or more bounding box coordinates, one per nodule detected | Sensitivity of 0.98 and false positives per image less than 1 | 2000 CT scans manually annotated by radiology resident and reviewed by thoracic radiologist. 200 CT scans annotated and adjudicated by three thoracic radiologists. |
| Lung Nodule Candidate False Positive Rejector ML | CNN with binary classification head | Previous output | Returns False if candidate appears to be a false positive, True otherwise. |
ROC >0.95 | The training data is taken from the output of lung nodule candidate generator item. |
| Lung Nodule Classifier ML | CNN with binary classification head | Previous output with True
|
Returns True if likely malignant. False if benign. |
ROC > 0.95 | 5000 bounding boxes of lung nodules. Annotated by a thoracic radiologist. |